Artykuł sponsorowany
Panele słoneczne a zmiany w przepisach dotyczących energetyki odnawialnej

W ostatnich latach coraz więcej przedsiębiorstw decyduje się na inwestycje w panele słoneczne. Ta decyzja wynika zarówno z rosnącej świadomości ekologicznej, jak i korzystnych zmian w przepisach dotyczących energetyki odnawialnej. W tym artykule przyjrzymy się wpływowi tych zmian na rozwój przedsiębiorstw oraz korzyściom płynącym z inwestycji w panele słoneczne.
Przeczytaj również: Krok po kroku: montaż blachodachówki na dachu w Toruniu
Przeczytaj również: Jak wybrać dobrego dewelopera w Gdyni: kluczowe kryteria wyboru
Zmiany w prawie a inwestycje w fotowoltaikę
Przeczytaj również: Różne rodzaje moskitier – które sprawdzą się w Twoim domu?
W ostatnich latach obserwujemy dynamiczny rozwój rynku fotowoltaicznego, który jest wspierany przez zmiany w przepisach dotyczących energetyki odnawialnej. Nowe regulacje wprowadzają m.in. możliwość sprzedaży nadwyżek wyprodukowanej energii do sieci oraz korzystniejsze warunki dla mikroinstalacji fotowoltaicznych. Dzięki temu przedsiębiorstwa mogą liczyć na dodatkowe źródło dochodu oraz szybszy zwrot z inwestycji w panele słoneczne.
Korzyści finansowe z inwestycji w panele słoneczne
Inwestycja w panele słoneczne to nie tylko odpowiedź na rosnące wymagania ekologiczne, ale także sposób na osiągnięcie korzyści finansowych. Przedsiębiorstwa, które zdecydują się na montaż instalacji fotowoltaicznej, mogą liczyć na obniżenie kosztów związanych z energią elektryczną oraz dodatkowe źródło dochodu ze sprzedaży nadwyżek energii. Ponadto, dzięki nowym przepisom, przedsiębiorstwa mogą korzystać z różnych form wsparcia finansowego, takich jak dotacje czy ulgi podatkowe.
Oszczędności energetyczne i ekologiczne aspekty inwestycji
Inwestycja w panele słoneczne przyczynia się również do zmniejszenia zużycia energii pochodzącej z konwencjonalnych źródeł, co przekłada się na niższe emisje dwutlenku węgla i innych szkodliwych substancji. Dzięki temu przedsiębiorstwa mogą poprawić swój wizerunek ekologiczny oraz spełniać coraz bardziej rygorystyczne normy środowiskowe. Ponadto, wykorzystanie energii słonecznej może wpłynąć na zmniejszenie zależności od niestabilnych cen energii na rynku oraz uniezależnienie się od dostawców energii.
Wnioski płynące z powyższych informacji pokazują, że inwestycja w panele słoneczne może być korzystna zarówno dla środowiska, jak i dla rozwoju przedsiębiorstw. Dzięki zmianom w przepisach dotyczących energetyki odnawialnej oraz rosnącej świadomości ekologicznej coraz więcej firm decyduje się na taki krok, przyczyniając się tym samym do rozwoju rynku fotowoltaicznego i poprawy stanu środowiska naturalnego.
Kategorie artykułów
Polecane artykuły
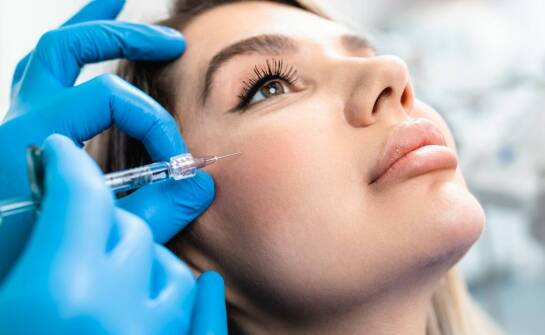
Korzyści z zastosowania polinukleotydów w zabiegach liftingujących.
Polinukleotydy to substancje stosowane w kosmetologii, które znajdują zastosowanie w zabiegach liftingujących. Dzięki właściwościom stymulującym regenerację tkanek mogą przyczynić się do poprawy elastyczności i nawilżenia skóry. Wprowadzenie do tematu pozwala zrozumieć, dlaczego te związki cieszą si

Jak mobilna fizjoterapia wspomaga procesy rehabilitacyjne po złamaniach?
Mobilna fizjoterapia z dojazdem Gorzów Wielkopolski jest wybierana m.in. w kontekście rehabilitacji po złamaniach. Elastyczność tej formy terapii pozwala dostosować sesje do potrzeb pacjentów, co może mieć znaczenie dla przebiegu postępowania rehabilitacyjnego. Terapeuci przyjeżdżają do domów, prowa